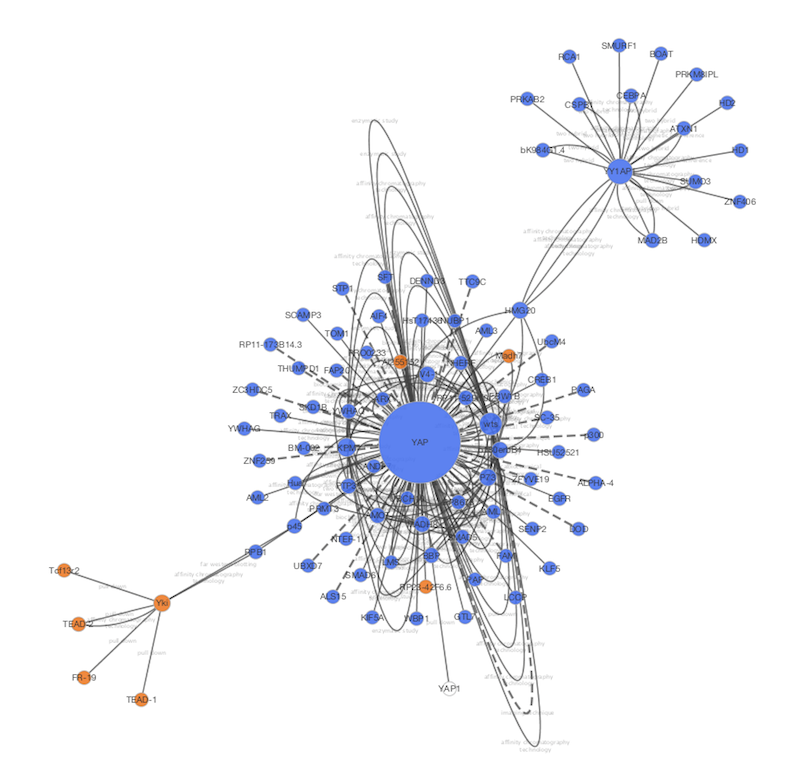

enhancedGraphics utilizes the baseĬustomGraphicsLayer for the two gradient types andįigure 1 shows examples of all of the gradients and charts that are provided by enhancedGraphics. PaintedShape which adds methods to return Shapes, Strokes, and Paints to draw arbitrary shapes. ImageCustomGraphicLayer, that adds a getPaint method that returns a suitable for painting an image on a node and (3) The API defines three types ofĬustomGraphicsLayers: (1) the base interface,ĬustomGraphicsLayers, that provides a getPaint method to return a simple for the node (2) EachĬ圜ustomGraphics object implements a getLayers method that generates the appropriate list ofĬustomGraphicLayers. TheĬ圜ustomGraphics object parses the String as appropriate. Internally, each enhancedGraphics chart type implements aĬ圜ustomGraphicsFactory that is registered with OSGiĬ圜ustomGraphicsFactory informs the visual mapping mechanism of the chart type (e.g.Ĭ圜ustomGraphics object given a String, which is the instruction column value. The drawing and display of the chart or graph is handled by enhancedGraphics methods that are called by the Cytoscape rendering engine. The format of the instruction column is type: arglist, where type is the type of gradient or chart, and arglist is a list of name=value pairs that specify the arguments to create the gradient or chart (see details and examples in the tables below). To use the gradients and charts provided by enhancedGraphics, an app or user would create two things: a column that contains the instructions for creating the chart, and a passthrough visual mapping that maps that column to one of the custom graphics visual properties. The intent is to provide a single, consistent, mechanism to draw charts and graphs on nodes as a general solution for diverse users and other app developers, mitigating the need to reinvent this capability in future apps.Īs part of the visual property mechanism, enhancedGraphics utilizes the Cytoscape 3 custom graphics API (.customgraphics). The mechanism supports saving and restoring charts, as well as high-quality image file output suitable for publication. For Cytoscape 3, we reimplemented this approach as an app to take advantage of the new architecture and custom graphics API. The initial version of this approach was developed for Cytoscape 2.8 as nodeCharts, which was used by clusterMakerħ, for example, and numerous users (see Figure 3 in the Jäger,Ĩ). We felt that a better approach would be to implement an app that provided general support for graphs and charts to users as well as to the developers of other apps. Each of these plugins and apps implemented their own graph and chart capabilities that are not accessible to other apps and in some cases not applicable outside of specific types of analyses. These include GOlorizeģ, which maps GO terms to pie charts on nodes GeneProĤ, which visualizes groups of nodes as pie charts VistaClaraĥ, which adds bar graphs to represent expression data and more recently MultiColoredNodesĦ. Over the years, there have been attempts to support more complex mappings of multiple data values onto node visuals in Cytoscape. 
This provides an extremely useful mapping between data values and a single visual property, but does not solve the need for more complex visualizations. 2 provides support for coloring and sizing nodes and node borders based on data values stored in the Node Table.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
